Protocol for use with NEBNext® Poly(A) mRNA Magnetic Isolation Module (NEB E7490) and NEBNext® Ultra RNA Library Prep Kit for Illumina® (E7530)
Symbols
 |
This is a point where you can safely stop the protocol and store the samples prior to proceeding to the next step in the protocol. |
 |
This caution sign signifies a step in the protocol that has two paths leading to the same end point but is dependent on a user variable, like the type of RNA input. |
 |
Colored bullets indicate the cap color of the reagent to be added |
The protocol has been optimized using high quality Universal Human Reference Total RNA. For PolyA mRNA selection, high quality RNA with RIN score > 7 (measured by Agilent Bioanalyzer®) is required.
RNA Sample Recommendations
The RNA sample should be free of salts (e.g., Mg2+, or guanidinium salts) or organics (e.g., phenol and ethanol).
Treat the RNA sample with DNase I to remove all traces of DNA. Remove DNase I after treatment.
AMPure XP Beads are required throughout the protocol. Allow beads to reach room temperature prior to use.
Starting Material: Total RNA (10 ng–1 μg) quantified by Bioanalyzer.
The protocol is optimized for approximately 200 bp RNA inserts. To generate libraries with longer RNA insert sizes, refer to Chapter 4 (Appendix A in the manual) for recommended fragmentation times and size selection conditions.
1.1.
 Preparation of First Strand Reaction Buffer and Random
Primer Mix
Preparation of First Strand Reaction Buffer and Random
Primer Mix 1.1.1 Prepare the First Strand Synthesis Reaction Buffer and Random Primer Mix as follows in a nuclease-free tube:
| COMPONENT | VOLUME |
|---|---|
 (pink) NEBNext First Strand Synthesis Reaction Buffer (pink) NEBNext First Strand Synthesis Reaction Buffer |
8 μl |
 (pink) NEBNext Random Primers (pink) NEBNext Random Primers |
2 μl |
| Nuclease-free water | 10 μl |
| Total Volume | 20 μl |
You can prepare the first strand synthesis reaction buffer later in the protocol, but it is important that it is ready before the elution in Step 1.2.37. The beads should not be allowed to dry out.
1.1.2 Mix thoroughly by pipetting up and down ten times.
Note: Keep the mix on ice.
1.2 mRNA Isolation, Fragmentation and Priming Starting with Total RNA
1.2.1. Dilute the total RNA with nuclease-free water to a final volume of 50 μl in a nuclease-free 0.2 ml PCR tube and keep on ice.
1.2.2. Aliquot 20 μl of NEBNext Oligo d(T)25 beads into a nuclease-free 0.2 ml PCR tube.
1.2.3. Wash the beads by adding 100 μl of RNA Binding Buffer (2X) to the beads. Pipette the entire volume up and down 6 times to mix thoroughly.
1.2.4. Place the tubes on the magnetic rack at room temperature for 2 minutes.
1.2.5. Remove and discard all of the supernatant from the tube. Take care not to disturb the beads.
1.2.6. Remove the tube from the magnetic rack.
1.2.7. Repeat steps 1.2.3. - 1.2.6.
1.2.8. Resuspend the beads in 50 μl of RNA Binding Buffer (2X) and add the 50 μl of total RNA sample from Step 1.2.1.
1.2.9. Place the tube on a thermal cycler and close the lid. Heat the sample at 65°C for 5 minutes and hold at 4°C to denature the RNA and facilitate binding of the poly-A mRNA to the beads.
1.2.10. Remove the tube from the thermal cycler when the temperature reaches 4°C.
1.2.11. Resuspend the beads. Pipette up and down slowly 6 times to mix thoroughly.
1.2.12. Place the tube on the bench and incubate at room temperature for 5 minutes to allow the mRNA to bind to the beads.
1.2.13. Resuspend the beads. Pipette up and down slowly 6 times to mix thoroughly.
1.2.14. Incubate for 5 minutes at room temperature to allow the RNA to bind to the beads.
1.2.15. Place the tube on the magnetic rack at room temperature for 2 minutes to separate the poly-A mRNA bound to the beads from the solution.
1.2.16. Remove and discard all of the supernatant. Take care not to disturb the beads.
1.2.17. Remove the tube from the magnetic rack.
1.2.18. Wash the beads by adding 200 μl of Wash Buffer to the tube to remove unbound RNA. Pipette the entire volume up and down 6 times to mix thoroughly.
1.2.19. Place the tube on the magnetic rack at room temperature for 2 minutes.
1.2.20. Remove and discard all of the supernatant from the tube. Take care not to disturb the beads.
1.2.21. Remove the tube from the magnetic rack.
1.2.22. Repeat steps 1.2.18. - 1.2.21.
1.2.23. Add 50 μl of Tris Buffer to each tube. Gently pipette the entire volume up and down 6 times to mix thoroughly.
1.2.24. Place the tube on the thermal cycler. Close the lid and heat the samples at 80°C for 2 minutes, then hold at 25°C to elute the Poly-A mRNA from the beads.
1.2.25. Remove the tube from the thermal cycler when the temperature reaches 25°C.
1.2.26. Add 50 μl of RNA Binding Buffer (2X) to the sample to allow the mRNA to re-bind to the beads. Gently pipette the entire volume up and down 6 times to mix thoroughly.
1.2.27. Incubate the tube at room temperature for 5 minutes.
1.2.28. Resuspend the beads. Pipette up and down slowly 6 times to mix thoroughly.
1.2.29. Incubate for 5 minutes at room temperature to allow the RNA to bind to the beads.
1.2.30. Place the tube on the magnetic rack at room temperature for 2 minutes.
1.2.31. Remove and discard all of the supernatant from the tube. Take care not to disturb the beads.
1.2.32. Remove the tubes from the magnetic rack.
1.2.33. Wash the beads by adding 200 μl of Wash Buffer. Gently pipette the entire volume up and down 6 times to mix thoroughly.
1.2.34. Place the tube on the magnetic rack at room temperature for 2 minutes.
1.2.35. Remove and discard all of the supernatant from the tube. Take care not to disturb the beads.
Note: It is important to remove all of the supernatant to successfully fragment the mRNA in the subsequent steps. Spin down the tube. Place the tube on the magnetic rack and with a 10 μl tip remove all of the wash buffer. Caution: Do not disturb beads that contain the mRNA.
1.2.36. Remove the tube from the magnetic rack.
Note: For RNA insert sizes > 200 nt, refer to Chapter 4 (Appendix A in the manual) for recommended fragmentation times.
1.2.37. Elute mRNA from the beads by adding 17 μl of the First Strand Synthesis Reaction Buffer and Random Primer mix prepared in Section 1.1.1. incubating the sample at 94°C for 15 minutes. Immediately, place the tubes on the magnetic rack.
1.2.38. Collect the purified mRNA by transferring 15 μl of the supernatant to a clean nuclease-free PCR Tube.
1.2.39. Place the tube on ice and proceed directly to first strand cDNA synthesis.
1.3 First Strand cDNA Synthesis
1.3.1. To the fragmented and primed mRNA (15 μl from above Section 1.2.38.) add the following components and mix by gentle pipetting:
| Component | Volume |
|---|---|
 (pink) Murine RNase Inhibitor (pink) Murine RNase Inhibitor |
0.5 μl |
 (pink) ProtoScript II Reverse Transcriptase (pink) ProtoScript II Reverse Transcriptase |
1 μl |
| Nuclease free water | 3.5 μl |
| Total Volume | 20 μl |
Note: If you are following recommendations in Appendix A in the manual, for longer RNA fragments, increase the incubation at 42°C from 15 minutes to 50 minutes in Step 1.3.2.
1.3.2.  Incubate the sample in a preheated thermal cycler as follows:
Incubate the sample in a preheated thermal cycler as follows:
10 minutes at 25°C
15 minutes at 42°C
15 minutes at 70°C
Hold at 4°C
1.3.3. Immediately, perform second strand cDNA synthesis reaction.
1.4 Perform Second Strand cDNA Synthesis
1.4.1. Add the following reagents to the First Strand Synthesis reaction (20 μl):
| Component | Volume |
|---|---|
 (orange) Second Strand Synthesis Reaction Buffer (orange) Second Strand Synthesis Reaction Buffer |
8 μl |
 (orange) Second Strand Synthesis Enzyme Mix (orange) Second Strand Synthesis Enzyme Mix |
4 μl |
| Nuclease free water | 48 μl |
| Total Volume | 80 μl |
1.4.2. Mix thoroughly by gentle pipetting.
1.4.3. Incubate in a thermal cycler for 1 hour at 16°C, with heated lid set at 40°C.
1.5 Purify the Double-stranded cDNA Using 1.8X Agencourt AMPure XP Beads
1.5.1. Vortex AMPure XP Beads to resuspend.
1.5.2. Add 144 μl (1.8X) of resuspended AMPure XP Beads to the second strand synthesis reaction (~80 μl). Mix well on a vortex mixer or by pipetting up and down at least 10 times.
1.5.3. Incubate for 5 minutes at room temperature.
1.5.4. Quickly spin the tube in a microcentrifuge to collect any sample on the sides of the tube. Place the tube on an appropriate magnetic rack to separate beads from supernatant. After the solution is clear (about 5 minutes), carefully remove and discard the supernatant. Be careful not to disturb the beads that contain DNA targets.
1.5.5. Add 200 μl of freshly prepared 80% ethanol to the tube while in the magnetic rack. Incubate at room temperature for 30 seconds, and then carefully remove and discard the supernatant.
1.5.6. Repeat Step 1.5.5. once for a total of 2 washing steps.
1.5.7 Air dry the beads for 5 minutes while the tube is on the magnetic rack with lid open.
1.5.8. Remove the tube from the magnet. Elute the DNA target from the beads into 60 μl 0.1X TE Buffer or 10 mM Tris-HCl pH 8.0. Mix well on a vortex mixer or by pipetting up and down. Quickly spin the tube and incubate for 2 minutes at room temperature. Place the tube in the magnetic rack until the solution is clear.
1.5.9. Remove 55.5 μl of the supernatant and transfer to a clean nuclease-free
PCR tube.
 Note: If you need to stop at this point in the protocol samples can be
stored at –20°C.
Note: If you need to stop at this point in the protocol samples can be
stored at –20°C.
1.6 Perform End Prep of cDNA Library
1.6.1. Mix the following components in a sterile nuclease-free tube:
| COMPONENT | VOLUME |
|---|---|
 (green) NEBNext End Repair Reaction Buffer (green) NEBNext End Repair Reaction Buffer |
6.5 μl |
 (green) NEBNext End Prep Enzyme Mix (green) NEBNext End Prep Enzyme Mix |
3 μl |
| Purified double stranded cDNA (Step 9, Section 1.5) | 55.5 μl |
| Total Volume | 65 μl |
1.6.2. Incubate the sample in a thermal cycler (with the heated lid set at 75°C) as follows:
30 minutes at 20°C
30 minutes at 65°C
Hold at 4°C
1.6.3. Proceed immediately to Adaptor Ligation.
1.7 Perform Adaptor Ligation
Dilute the  (red) NEBNext Adaptor* for Illumina (15 μM) to 1.5 μM with a 10-fold dilution (1:9) with 10 mM Tris-HCl and 10 mM NaCl for immediate use.
(red) NEBNext Adaptor* for Illumina (15 μM) to 1.5 μM with a 10-fold dilution (1:9) with 10 mM Tris-HCl and 10 mM NaCl for immediate use.
1.7.1. Add the following components directly to the prep reaction mixture. (Caution: do not pre-mix the components to prevent adaptor-dimer formation):
| COMPONENT | VOLUME |
|---|---|
| End Prep Reaction | 65 μl |
 (red) Blunt/TA Ligase Master Mix (red) Blunt/TA Ligase Master Mix |
15 μl |
| Diluted NEBNext Adaptor* | 1 μl |
| Nuclease-free Water | 2.5 μl |
| Total Volume | 83.5 μl |
*The adaptor is provided in NEBNext Singleplex (NEB #E7350) or NEBNext Multiplex (NEB #E7335, #E7500, #E6609, #E7710, #E7730 or #E7600) Oligos for Illumina.
1.7.2. Mix by pipetting followed by a quick spin to collect all liquid from the sides of the tube.
1.7.3. Incubate 15 minutes at 20°C in a thermal cycler. Turn off the heated lid on the thermal cycler.
1.7.4. Add 3 μl of  (red) USER Enzyme to the ligation mixture from Step 1.7.3. Mix well and incubate at 37°C for 15 minutes.
(red) USER Enzyme to the ligation mixture from Step 1.7.3. Mix well and incubate at 37°C for 15 minutes.
Note: This step is only required for use with NEBNext Adaptors. USER Enzyme can be found in the NEBNext SinglePlex (NEB #E7350) or Multiplex (NEB #E7335, #E7500, #E7710, #E7730, #E6609, #E7600) Oligos for Illumina.
 A precipitate can form upon thawing of the NEBNext Q5 Hot Start HiFi PCR Master Mix. To ensure optimal performance, place the master mix at room temperature while purifying the ligation reaction. Once thawed, gently mix by inverting the tube several times.
A precipitate can form upon thawing of the NEBNext Q5 Hot Start HiFi PCR Master Mix. To ensure optimal performance, place the master mix at room temperature while purifying the ligation reaction. Once thawed, gently mix by inverting the tube several times.
1.8 Purify the Ligation Reaction Using AMPure XP Beads
 Note: If you are selecting for larger size fragments (> 200 nt) follow the size selection recommendations in Appendix A, Chapter 4 in the manual.
Note: If you are selecting for larger size fragments (> 200 nt) follow the size selection recommendations in Appendix A, Chapter 4 in the manual.
1.8.1 Add nuclease-free water to the ligation reaction to bring the reaction volume to 100 μl.
Note: X refers to the original sample volume of 100 μl from the above step.
1.8.2. Add 100 μl (1.0X) resuspended AMPure XP Beads and mix well on a vortex mixer or by pipetting up and down at least 10 times.
1.8.3. Incubate for 5 minutes at room temperature.
1.8.4. Quickly spin the tube in a microcentrifuge and place the tube on an appropriate magnetic rack to separate beads from the supernatant. After the solution is clear (about 5 minutes), discard the supernatant that contain unwanted fragments (Caution: do not discard the beads).
1.8.5. Add 200 μl of freshly prepared 80% ethanol to the tube while in the magnetic rack. Incubate at room temperature for 30 seconds, and then carefully remove and discard the supernatant.
1.8.6. Repeat Step 1.8.5. once for a total of 2 washing steps.
1.8.7. Briefly spin the tube, and put the tube back in the magnetic rack.
1.8.8. Completely remove the residual ethanol, and air dry beads for 5 minutes while the tube is on the magnetic rack with the lid open.
Caution: Do not overdry the beads. This may result in lower recovery of DNA target.
1.8.9. Remove the tube from the rack. Elute DNA target from the beads with 52 μl 0.1X TE or 10 mM Tris-HCI. Mix well on a vortex mixer or by pipetting up and down, and incubate for 2 minutes at room temperature. Put the tube in the magnetic rack until the solution is clear.
1.8.10. Transfer 50 μl supernatant to a clean PCR tube. Discard beads.
1.8.11. To the 50 μl supernatant, add 50 μl (1.0X) of the resuspended AMPure XP Beads and mix well on a vortex mixer or by pipetting up and down at least 10 times.
1.8.12. Incubate for 5 minutes at room temperature.
1.8.13. Quickly spin the tube in a microcentrifuge and place the tube on an appropriate magnetic rack to separate beads from the supernatant. After the solution is clear (about 5 minutes), discard the supernatant that contains unwanted fragments (Caution: do not discard the beads).
1.8.14. Add 200 μl of freshly prepared 80% ethanol to the tube while in the magnetic rack. Incubate at room temperature for 30 seconds, and then carefully remove and discard the supernatant.
1.8.15. Repeat Step 1.8.14. once for a total of 2 washing steps.
1.8.16. Briefly spin the tube, and put the tube back in the magnetic rack.
1.8.17. Completely remove the residual ethanol, and air dry beads for 5 minutes while the tube is on the magnetic rack with the lid open.
1.8.18. Remove the tube from the rack. Elute DNA targtet from the beads with 22 μl 0.1X TE or 10 mM Tris-HCl. Mix well on a vortex mixer or by pipetting up and down, and incubate for 2 minutes at room temperature. Put the tube in the magnetic rack until the solution is clear.
1.8.19. Without disturbing the bead pellet, transfer 20 μl of the supernatant to a clean PCR tube and proceed to PCR enrichment.
1.9 PCR Enrichment of Adaptor Ligated DNA
 Follow Section 1.9A if you are using the following oligos (10 μM primer):
Follow Section 1.9A if you are using the following oligos (10 μM primer):
NEBNext Singleplex Oligos for Illumina (NEB #E7350)
NEBNext Multiplex Oligos for Illumina (Set 1, NEB #E7335)
NEBNext Multiplex Oligos for Illumina (Set 2, NEB #E7500)
NEBNext Multiplex Oligos for Illumina (Set 3, NEB #E7710)
NEBNext Multiplex Oligos for Illumina (Set 4, NEB #E7730)
NEBNext Multiplex Oligos for Illumina (Dual Index Primers,
NEB #E7600)
Follow Section 1.9B if you are using NEBNext Multiplex Oligos for Illumina (96 Index Primers, NEB #E6609)
1.9A PCR Library Enrichment
1.9A.1. To the cDNA (20 μl) from Step 1.8.19. add the following components and mix by gentle pipetting:
| COMPONENT | VOLUME |
|---|---|
 (blue) NEBNext Q5 Hot Start HiFi PCR Master Mix (blue) NEBNext Q5 Hot Start HiFi PCR Master Mix |
25 μl |
 (blue) Index Primer/i7 Primer*,** (blue) Index Primer/i7 Primer*,** |
2.5 μl |
 (blue) Universal PCR Primer/i5 Primer*, *** (blue) Universal PCR Primer/i5 Primer*, *** |
2.5 μl |
| Total Volume | 50 μl |
* The primers are provided in NEBNext Singleplex (NEB #E7350) or Multiplex (NEB #E7335, #E7500, #E7710, #E7730, #E7600) Oligos for Illumina. For use with Dual Index Primers (NEB #E7600), look at the NEB #E7600 manual for valid barcode combinations and tips for setting up PCR reactions.
** For use with NEBNext Multiplex Oligos (NEB #E7335, #E7500, #E7710, #E7730) use only one Index Primer per PCR reaction. For use with Dual Index Primers (NEB #E7600) use only one i7 Primer per reaction.
*** For use with Dual Index Primers (NEB #E7600) use only one i5 Primer per reaction.
1.9A.2. PCR Cycling Conditions
| CYCLE STEP | TEMPERATURE | TIME | CYCLES |
|---|---|---|---|
| Initial Denaturation | 98°C | 30 seconds | 1 |
| Denaturation Annealing/Extension |
98°C 65°C |
10 seconds 75 seconds |
12–15*, ** |
| Final Extension | 65°C | 5 minutes | 1 |
| Hold | 4°C | ∞ |
* The number of PCR cycles should be adjusted based on RNA input. If 10 ng total RNA is the starting input, it is recommended to perform 15 cycles of PCR. However, optimization of PCR cycle number may be required to avoid-amplification.
** It is important to limit the number of PCR cycles to avoid overamplification.
If overamplification occurs, larger molecular weight products (> 500 bp) will appear on the Bioanalyzer trace.
1.9A.3. Proceed to Section 1.10 (Purify the PCR Reaction using Agencourt AMPure XP Beads).
1.9B PCR Library Enrichment
1.9B.1. To the cDNA (20 μl) from Step 1.8.19. add the following components and mix by gentle pipetting:
| COMPONENT | VOLUME |
|---|---|
 (blue) Index/ Universal Primer Mix* (blue) Index/ Universal Primer Mix* |
5 μl |
 (blue) NEBNext Q5 Hot Start HiFi PCR Master Mix (blue) NEBNext Q5 Hot Start HiFi PCR Master Mix |
25 μl |
| Total Volume | 50 μl |
* The primers are provided in NEBNext Multiplex Oligos for Illumina, NEB #E6609. Refer to NEB #E6609 manual for valid barcode combinations and tips for setting up PCR reactions.
1.9B.2. PCR Cycling Conditions
| CYCLE STEP | TEMPERATURE | TIME | CYCLES |
|---|---|---|---|
| Initial Denaturation | 98°C | 30 seconds | 1 |
| Denaturation Annealing/Extension |
98°C 65°C |
10 seconds 75 seconds |
12–15*, ** |
| Final Extension | 65°C | 5 minutes | 1 |
| Hold | 4°C | ∞ |
* The number of PCR cycles should be adjusted based on RNA input. If 10 ng total RNA is the starting input, it is recommended to perform 15 cycles of PCR. However, optimization of PCR cycle number may be required to avoid-amplification.
** It is important to limit the number of PCR cycles to avoid overamplification. If overamplification occurs, larger molecular weight products (> 500 bp) will appear on the Bioanalyzer trace.
1.9B.3. Proceed to Section 1.10 (Purify the PCR Reaction using Agencourt AMPure XP Beads).
1.10 Purify the PCR Reaction using Agencourt AMPure XP Beads
Note: X refers to the original sample volume from the above step.
1.10.1. Vortex Agencourt AMPure XP Beads to resuspend.
1.10.2. Add 45 μl (0.9X) of resuspended Agencourt AMPure XP Beads to the PCR reaction (~ 50 μl). Mix well on a vortex mixer or by pipetting up and down at least 10 times.
1.10.3. Incubate for 5 minutes at room temperature.
1.10.4. Quickly spin the tube in a microcentrifuge and place the tube on an appropriate magnetic rack to separate beads from the supernatant. After the solution is clear (about 5 minutes), carefully remove and discard the supernatant. Be careful not to disturb the beads that contain DNA targets.
1.10.5. Add 200 μl of freshly prepared 80% ethanol to the tube while in the magnetic rack. Incubate at room temperature for 30 seconds, and then carefully remove and discard the supernatant.
1.10.6. Repeat Step 1.10.5. once for a total of 2 washing steps.
1.10.7. Air dry the beads for 5 minutes while the tube is on the magnetic rack with the lid open.
1.10.8. Remove the tube from the rack. Elute the DNA target from the beads into 23 μl 0.1X TE. Mix well on a vortex mixer or by pipetting up and down, quickly spin the tube in a microcentrifuge and incubate for 2 minutes at room temperature. Place it in the magnetic rack until the solution is clear.
1.10.9. Transfer 20 μl of the supernatant to a clean PCR tube, and store at –20°C.
1.11 Assess library quality on a Bioanalyzer® (Agilent High Sensitivity Chip).
1.11.1. Dilute 2–3 μl of the library in 10 mM Tris or 0.1X TE.
1.11.2. Run 1 μl in a DNA High Sensitivity Chip.
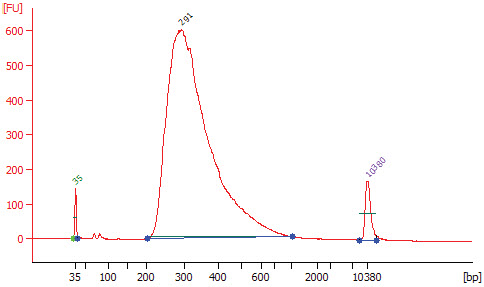
1.11.3. Check that the electropherogram shows a narrow distribution with a peak size approximately 300 bp. Note: If a peak at ~ 80 bp (primers) or 128 bp (adaptor-dimer) is shown in the Bioanalyzer traces, bring up the sample volume (Step 1.10.9.) to 50 μl exactly with nuclease-free water and repeat the AMPure XP Bead clean up step (Section 1.10).