Time-Saver™ Qualified Restriction Enzymes
Choose Type:
- Do degenerate recognition sites need to be palindromic?
- How can I generate a restriction enzyme site map for my sequence?
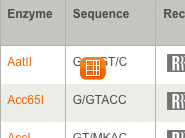
- How can I search for a restriction enzyme by sequence, overhang or name?
- How should I stop my restriction digest?
- How stable is a particular restriction enzyme?
- I don't see any cleavage after my restriction digest. What factors can interfere with cleavage?
- Is extended digestion (incubation times > 1 hour) recommended?
- What does it mean to be Time-Saver™ qualified?
- What information is available in the Restriction Enzyme Database (REBASE)?
- When is star activity a concern?
- When should I choose the HF version of the enzyme?
- How should I set up a restriction digest?
- What does HF® refer to following the name of a restriction enzyme?
-
Restriction Enzymes at NEB: Over 30 years of Innovation
-
A Modern Day Gene Genie Sir Richard Roberts on Rebase
- DNA Sequences and Maps Tool
- Alphabetized List of Recognition Sequences
- Cleavage Of Supercoiled DNA
- Compatible Cohesive Ends and Generation of New Restriction Sites
- Dam-Dcm and CpG Methylation
- Enzymes with Multiple Recognition Sequences
- Enzymes with Nonpalindromic Sequences
- Frequencies of Restriction Sites
- Interrupted Palindromes
- Isoschizomers
- Recleavable Blunt Ends
- Recleavable Filled-in 5' Overhangs
- Restriction Enzyme Troubleshooting Guide
- Activity at 37°C for Restriction Enzymes with Alternate Incubation Temperatures
- Activity of Restriction Enzymes in PCR Buffers
- Alteration of Apparent Recognition Specificities Using Methylases
- Cleavage Close to the End of DNA Fragments
- Dam and Dcm Methylases of E. coli
- Double Digests
- Heat Inactivation
- NEBuffer Activity/Performance Chart with Restriction Enzymes
- Optimizing Restriction Endonuclease Reactions
- Restriction Enzyme Diluent Buffer Compatibility
- Restriction Enzyme Tips
- Restriction of Foreign DNA by E. coli K-12
- Site Preferences
- Star Activity
Feature Articles
Web Tools
Selection Tools
Troubleshooting Guides
Usage Guidelines
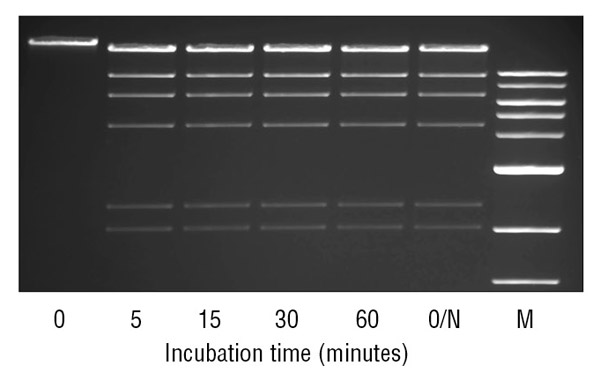
- Enzymes powerful enough to digest in 5-15 minutes
- Flexible enough for overnight incubation
- No special formulation
- No added expense
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.