RNA Cleanup
Choose Type:
- Can I purchase Monarch® buffers and columns separately?
- Are the columns in the Monarch RNA Cleanup Kits the same as those in the Monarch Total RNA Miniprep Kit (NEB #T2010)?
- Can I get better recovery with the Monarch RNA Cleanup Kits if I do a second elution with my eluent from the first elution?
- Can I use the Monarch RNA Cleanup Kit to cleanup up my DNase I-treated RNA?
- Can I use the Monarch RNA Cleanup Kits to cleanup my in vitro transcription (IVT) reaction?
- Can I use the Monarch RNA Cleanup Kits to cleanup RNA after a TRIzol®/chloroform extraction?
- Can I use the Monarch RNA Cleanup Kits to purify RNA from agarose gels?
- Do you have a protocol for separating small and large RNAs into separate fractions?
- How can I assess RNA integrity and purity?
- Is the Monarch RNA Cleanup Kit (NEB #T2040) compatible with the EnGen sgRNA Synthesis Kit, S. pyogenes?
- What factors affect my (A260/A230) when using the Monarch RNA Cleanup Kits?
- What is the composition of each buffer provided with the Monarch RNA Cleanup Kits?
- What is the maximum binding capacity of the Monarch RNA Cleanup Column provided with the Monarch RNA Cleanup Kit?
- What is the smallest volume of nuclease-free water that can be used for elution with the Monarch RNA Cleanup Columns?
- What size RNA can be purified with the Monarch RNA Cleanup Kit?
- Are the Monarch RNA Cleanup Kits compatible with Luna RT-qPCR reagents?
- Are the Monarch RNA Cleanup Kits compatible with NEBNext reagents for RNA library prep?
- My sample turned cloudy after adding the Monarch RNA Cleanup Binding Buffer and ethanol. Is this normal?
- Why do I need to incubate my column for 5 minutes with elution buffer (nuclease-free water)?
- Can I do an on-column DNase I treatment with the Monarch RNA Cleanup Columns?
- Monarch® RNA Cleanup Kit Protocol
- Purification of RNA from the Aqueous Phase Following TRIzol®/Chloroform Extraction using the Monarch® RNA Cleanup Kits
- Extraction of RNA from Agarose Gels using the Monarch® RNA Cleanup Kits
- Separation of Large and Small RNA into Fractions using the Monarch® RNA Cleanup Kits
- RNA Reaction Cleanup using the Monarch Total RNA Miniprep Kit (NEB #T2010)
- Monarch Nucleic Acid Purification Brochure
- RNA Metro Map Poster
- RNA Synthesis Brochure
- RNA Technical Guide
- Troubleshooting Guide for RNA Cleanup
- Guidelines for RNA Quantitation
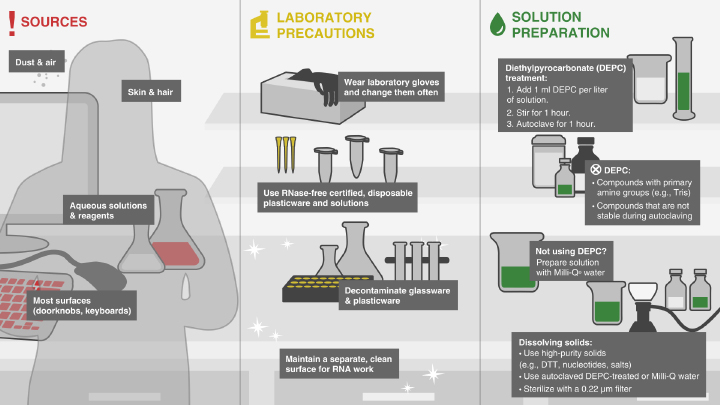
- Guidelines for Working with RNA During RNA Cleanup
Brochures
Troubleshooting Guides
Usage Guidelines
| Applications | |
|---|---|
|
RNA Cleanup and Concentration (including from the TRIzol aqueous phase) |
RNA purified by other methods can be further purified |
| Enzymatic Reaction Cleanup | Enzymes such as RNA polymerases, DNase I, Proteinase K and phosphatases are removed allowing efficient desalting |
| In vitro Transcription Cleanup | Enzymes and excess NTPs are removed to yield highly pure synthesized RNA |
| RNA Gel Extraction | Purification of RNA from agarose gels |
| RNA Fractionation | Fractionation of RNA into small and large RNA pools |
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. The use of this product may require the buyer to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
This product is intended for research purposes only. This product is not intended to be used for therapeutic or diagnostic purposes in humans or animals.